Journal Description
Microbiology Research
Microbiology Research
is an international, scientific, peer-reviewed open access journal published quarterly online by MDPI (from Volume 11 Issue 2-2020).
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), Embase, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.6 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
Impact Factor:
1.5 (2022);
5-Year Impact Factor:
1.6 (2022)
Latest Articles
Urinary Tract Infections in a Single Hospital in Central Portugal, a 5-Year Analysis
Microbiol. Res. 2024, 15(2), 850-863; https://doi.org/10.3390/microbiolres15020055 - 17 May 2024
Abstract
►
Show Figures
Urinary tract infections are defined as the presence of microorganisms in any part of the urinary system, with the exception of the distal urethra. A majority of them are uncomplicated infections that are resolved on an outpatient basis, with empirical therapy. The objectives
[...] Read more.
Urinary tract infections are defined as the presence of microorganisms in any part of the urinary system, with the exception of the distal urethra. A majority of them are uncomplicated infections that are resolved on an outpatient basis, with empirical therapy. The objectives of this work were to study the sociodemographic characteristics of patients, analyze associated strains and examine the response of the main microorganisms to antibiotics. A retrospective observational study of all positive urine cultures between 2018 and 2022 was carried out at an institution (8340 samples). Sociodemographic data were also collected. In total, 61.3% were women, with an average age of 63.4 years, and 43.2% were from the Emergency Department. A total of 13.5% were fitted, 56% of whom were women. Also, 95.9% were not taking any antibiotics, and among the individuals who were taking antibiotics, 50% were injected. Escherichia coli (53.5%) and Klebsiella pneumoniae (13.8%) are identified as the most prevalent strains. In the time periods analyzed, Escherichia coli decreased its resistance to 11 antibiotics and increased to 5 antibiotics, while Klebsiella pneumoniae decreased to 7 and increased to 7, with emphasis on the presence of 3 antibiotics with a resistance rate of 100% to all Klebsiella pneumoniae strains identified in 2022.
Full article
Open AccessCommunication
Molecular Analysis of the Microbial Guild Fixing Nitrogen in Ricefield Soils in Missouri
by
Prithi R. Sawli, Mark A. Buchheim and Mark A. Schneegurt
Microbiol. Res. 2024, 15(2), 841-849; https://doi.org/10.3390/microbiolres15020054 - 17 May 2024
Abstract
►▼
Show Figures
Non-symbiotic diazotrophic microbes are important contributors to global N budgets in cereal crops. Knowledge of the biogeography of the organisms in this functional guild increases our understanding of biological N fixation in diverse locations and climates. Here, we describe the diazotrophic community in
[...] Read more.
Non-symbiotic diazotrophic microbes are important contributors to global N budgets in cereal crops. Knowledge of the biogeography of the organisms in this functional guild increases our understanding of biological N fixation in diverse locations and climates. Here, we describe the diazotrophic community in the previously unstudied, extensive ricefields of southeast Missouri, using restriction fragment length polymorphism (RFLP) analysis and sequencing of nifH gene clones. While nine RFLP patterns were observed in random nifH clones, these groups were not all supported by gene sequencing, suggesting that the RFLP of nifH genes alone is not suitable for describing diazotrophic guilds. Dozens of nifH clones from Missouri ricefield soils were sequenced and analyzed phylogenetically. The nifH genes detected were predominantly from Geobacteraceae, most closely related to Geobacter and Geomonas species. There were substantial clusters of nifH clones most closely related to Desulfovibrionales and other Proteobacteria. Many of the clones did not closely cluster with nifH sequences from known isolates or clades. No cyanobacterial or archaeal sequences were detected in the Missouri ricefield soils. The microbial guild fixing N appeared to be rich in anaerobes and lithotrophs. Organisms in Geobacter and Geomonas seem to be cosmopolitan, but endemism was evident, since nifH clones were recovered that formed clusters not previously reported from ricefields in other locations.
Full article

Figure 1
Open AccessArticle
Endophytic Fungal Diversity in Hardwickia binata: Bridging the Gap between Traditional and Modern Techniques
by
Michael Joe Xavier Sneha, Myithili Thangavel, Israel Mani, Pandy Rajapriya, Nagendraprabhu Ponnuraj and Mohan Pandi
Microbiol. Res. 2024, 15(2), 823-840; https://doi.org/10.3390/microbiolres15020053 - 16 May 2024
Abstract
►▼
Show Figures
Endophytic fungus is crucial for maintaining plant health and defense mechanisms, acting as protective barriers against pathogens, and producing medicinally beneficial bioactive compounds. Genome sequencing and metagenomics have significantly enhanced the understanding of fungal diversity and metabolic capabilities, enabling the identification of new
[...] Read more.
Endophytic fungus is crucial for maintaining plant health and defense mechanisms, acting as protective barriers against pathogens, and producing medicinally beneficial bioactive compounds. Genome sequencing and metagenomics have significantly enhanced the understanding of fungal diversity and metabolic capabilities, enabling the identification of new genes and substances. Traditional culture-dependent methods have been complemented by culture-independent techniques, offering a more comprehensive view of fungal diversity. Using both culture-dependent and culture-independent techniques, the present research investigation explored the diversity of endophytic fungi encountered in the foliage of Hardwickia binata. The study examined the topographical characteristics and nutritional content of soil samples collected from the locality of the selected plant sample, H. binata, to better comprehend the effects on the plant’s growth. The balanced nutrient constituted approximately a pH of 7.2, which suggested an alkaline nature and promoted plant development. The ratio of nitrogen, phosphorous, and potassium remained 3:1:1. A total of 25 fungal isolates, categorized into 17 morphotypes, were obtained using the culture-dependent approach; Curvularia and Nigrospora emerged as the most common genera. Furthermore, the prediction of the ITS2 secondary structure supports the identification of species, highlighting a wide variety of fungal species present in H. binata. The culture-independent approach generated 69,570 high-quality sequences, identifying 269 Operational Taxonomic Units (OTUs). The dominant Ascomycota phylum, along with various genera, indicated a rich fungal community associated with H. binata. This study advances the understanding of the endophytic fungus communities that are associated with H. binata and the nature of soil ecology. The findings emphasize the significance of holistic techniques in the study of microbial dynamics within plant systems as well as their implications for ecosystem management and plant health.
Full article

Figure 1
Open AccessReview
Microbial Diversity and Nitrogen Cycling in Peat and Marine Soils: A Review
by
Akshatha Soratur, Balu Alagar Venmathi Maran, Ahmad Syazni Kamarudin and Kenneth Francis Rodrigues
Microbiol. Res. 2024, 15(2), 806-822; https://doi.org/10.3390/microbiolres15020052 - 14 May 2024
Abstract
Nitrogen is an essential nutrient for living organisms in peat and marine soils, and its transformation within the soil matrix is a complex process mediated by various microbes that inhabit these ecological niches. The metabolism of nitrogen is governed by microbially mediated biogeochemical
[...] Read more.
Nitrogen is an essential nutrient for living organisms in peat and marine soils, and its transformation within the soil matrix is a complex process mediated by various microbes that inhabit these ecological niches. The metabolism of nitrogen is governed by microbially mediated biogeochemical transformations, such as nitrification, anammox, and denitrification, which contribute to the assimilated pool of nitrogen and fixed nitrogen loss. One of the major challenges facing the field of peat and marine microbiology is the lack of understanding of the correlation between ecosystem-driven nitrogen transformation and microbial diversity. This is crucial because of growing concerns regarding the impacts of human-induced activities and global climate change on microbial nitrogen-cycling processes in peat and marine soils. Thus, this review aimed to provide a comprehensive overview of the current understanding of the microbial communities involved in peat and marine nitrification, anammox, and denitrification; the factors influencing the niche differentiation and distribution of the main functional components; the genes involved; and the main effects of human-induced activities and global climate change on the peat and marine nitrogen cycle. The implications of this review will facilitate an understanding of the complex mechanisms associated with ecosystem function in relation to nitrogen cycling, the role of peat and marine soils as carbon sinks, pollution remediation using naturally occurring populations of diverse microbes, and the development of policies to mitigate the effects of anthropogenic influences in peat and marine soils.
Full article
(This article belongs to the Special Issue Microorganisms as a Tool for Restoring the Environment)
►▼
Show Figures

Figure 1
Open AccessArticle
In Vitro Evaluation of Gastrointestinal Stability of Pediococcus pentosaceus Isolated from Fermented Maize and Pearl Millet for Possible Novel Chicken Probiotic Development
by
Gifty Ziema Bumbie, Leonardo Abormegah, Peter Asiedu, Akua Durowaa Oduro-Owusu, Kwame Owusu Amoah, Frederick Danso, Bernard Bortei Bortieh, Theresah Nkrumah, Taha Mohamed Mohamed and Zhiru Tang
Microbiol. Res. 2024, 15(2), 787-805; https://doi.org/10.3390/microbiolres15020051 - 14 May 2024
Abstract
Research has identified certain bio-based products, such as probiotics, as alternatives to antibiotics for use in animal feed. They are capable of controlling, preventing or minimizing the colonization of the gastrointestinal tract by pathogenic bacteria. To isolate Pediococcus spp. and assess its technological
[...] Read more.
Research has identified certain bio-based products, such as probiotics, as alternatives to antibiotics for use in animal feed. They are capable of controlling, preventing or minimizing the colonization of the gastrointestinal tract by pathogenic bacteria. To isolate Pediococcus spp. and assess its technological properties for possible probiotic development, maize and pearl millet were used. The cereals were steeped and wet milled after 48 h of fermentation. The milled cereals were kneaded into dough for 24 h, after which a 10% slurry was prepared for tenfold serial dilution to enumerate the LAB by employing pour plate techniques using MRS Agar. Based on the cell morphology of the isolated bacteria, eight isolates (four from maize and four from millet) that were selected for identification using MALDI-TOF MS showed that five were Pediococcus pentosaceus (P. pentosaceus), one was Pediococcus acidilactici, and two did not match any organism. Subsequently, the six isolates were labeled as MZ1, MZ2, MZ3, MZ4 for the maize isolate and MLT5 and MLT7 for the millet isolate. The six Pediococcus spp. were assessed in vitro for acid and bile salt tolerance, gastric juice and intestinal fluid tolerance and antibiotic resistance and antimicrobial susceptibility tests were performed, and the survivability rate of the strains was calculated. With regard to the mean count, there was a reduction in log10 CFU/mL under the lower pH conditions and their duration of exposure with regard to time. Among the isolates, no differences were noted at the various periods of exposure (0 h, 1 h, 2 h and 3 h) at pH 4 (p > 0.05). However, significant differences were noted at pH 3, 2 and 1 among the isolates (p < 0.05). The percentage survival of MZ4 and MLT7 at pH 1 was higher compared to the other isolates at 0 h. Significant differences were observed among the isolated at pH 2, 3 and 4 across the various periods. The mean count of the isolates in gastric juice was similar at 0 and 1 h, but significant differences were noted at 2 and 3 h, where MLT7 was highest (p < 0.05). A similar trend was observed for percentage survival. The mean count and the percentage survival of isolates under different concentrations of bile salt were similar. Significant differences were noted among isolates in both mean count and percentage survival when exposed to intestinal fluid (p < 0.05). All of the isolates were highly tolerant to the antibiotics tested and possessed antibacterial properties against the selected pathogens. The LABs proved to be good probiotic materials, according to the results obtained. However, the Pediococcus strain MLT7 proved to be the LAB of choice; therefore, its molecular identity was verified using the 16S rRNA sequence and was labeled as Pediococcus pentosaceus GT001 after it was discovered to have 100% similarity with some strains of Pediococcus pentosaceus.
Full article
Open AccessArticle
Exploring the Antimicrobial and Antioxidant Activities of Streptomyces sp. EIZ2 Isolated from Moroccan Agricultural Soil
by
Said Rammali, Fatima Zahra Kamal, Mohamed El Aalaoui, Abdellatif Rahim, Aziz Baidani, Khadija Dari, Abdelkrim Khattabi, Alin Ciobică, Bogdan Novac, Antoneta Petroaie, Radu Lefter and Bouchaib Bencharki
Microbiol. Res. 2024, 15(2), 762-786; https://doi.org/10.3390/microbiolres15020050 - 13 May 2024
Abstract
►▼
Show Figures
Antibiotics play a crucial role in preserving and improving public health, saving millions of lives every year. However, their effectiveness is currently under threat due to the ability of bacteria to adapt and develop resistance to these treatments. Therefore, this study was carried
[...] Read more.
Antibiotics play a crucial role in preserving and improving public health, saving millions of lives every year. However, their effectiveness is currently under threat due to the ability of bacteria to adapt and develop resistance to these treatments. Therefore, this study was carried out on two soil samples collected in two areas of Arba Aounate, Sidi Bennour province, Morocco, to identify natural antibiotic-producing Actinobacteria capable of combating multi-drug-resistant (MDR) bacteria. A primary screening revealed that of the 50 isolates, 16 exhibited antimicrobial activity against Pseudomonas aeruginosa ATCC 27,853, Staphylococcus aureus ATCC 25,923, Escherichia coli ATCC 25,922, and Candida albicans ATTC 60193. A secondary screening showed that of the 16 isolates, only EIZ1 and EIZ2 isolates displayed outstanding antimicrobial and antifungal activity against 6 MDR bacteria, including Escherichia coli 19L2418, Listeria monocytogenes, Proteus sp. 19K1313, Klebsiella pneumoniae 19K 929, Proteus vulgaris 16C1737, and Klebsiella pneumoniae 20B1572. These two isolates were also characterized culturally, morphologically, physiologically, and biochemically. Afterward, the amplification of 16S rRNA revealed that isolate EIZ2 was 99.06% strongly related to the genus Streptomyces. Furthermore, this extract exhibits strong antioxidant activity against DPPH and ABTS free radicals and elevated ferric-reducing antioxidant power. A significant (p < 0.0001) positive correlation was observed between antioxidant activities and total phenolic and flavonoid contents. A GC–MS analysis of the ethyl acetate extract revealed the presence of 10 compounds, mainly diethyl phthalate (97%) and benzeneacetic acid (94%). This research demonstrates that Streptomyces sp. strain EIZ2 represents a potential source of antimicrobial and antioxidant compounds. These compounds could offer considerable potential as therapeutic agents, paving the way for future developments in medical applications.
Full article

Figure 1
Open AccessArticle
A Quadruplex Reverse Transcription Quantitative Polymerase Chain Reaction for Detecting Canine Coronavirus, Canine Rotavirus, Canine Parvovirus, and Canine Distemper Virus
by
Yandi Shi, Feng Long, Kaichuang Shi, Mengyi He, Yuwen Shi, Shuping Feng, Yanwen Yin, Xiankai Wei and Zongqiang Li
Microbiol. Res. 2024, 15(2), 746-761; https://doi.org/10.3390/microbiolres15020049 - 10 May 2024
Abstract
Background: Canine coronavirus (CCoV), canine rotavirus (CRV), canine parvovirus (CPV), and canine distemper virus (CDV) cause gastroenteritis in dogs, and co-infections of these pathogens are common in China. In particular, CCoV and CRV are confirmed to have important zoonotic potential and cause public
[...] Read more.
Background: Canine coronavirus (CCoV), canine rotavirus (CRV), canine parvovirus (CPV), and canine distemper virus (CDV) cause gastroenteritis in dogs, and co-infections of these pathogens are common in China. In particular, CCoV and CRV are confirmed to have important zoonotic potential and cause public health issues. It is difficult to diagnose these diseases based only on clinical manifestations and pathological damage. Methods: In this study, four pairs of specific primers and probes targeting the CCoV M, CRV VP7, CPV VP2, and CDV N genes were designed. The reaction conditions, including the primer and probe concentrations, annealing temperatures, and reaction cycles, were optimized for the development of a quadruplex RT-qPCR for the detection of CCoV, CRV, CPV, and CDV. The assay was used to test 1028 clinical samples to validate its application. Results: A quadruplex RT-qPCR was successfully established for the differential detection of CCoV, CRV, CPV, and CDV, with good specificity, high sensitivity, and excellent repeatability. The assay could specifically detect CCoV, CRV, CPV, and CDV without cross-reactivity with the other canine viruses tested. It showed high sensitivity with limits of detection (LOD) of 1.1 × 102 copies/reaction for all four plasmid constructs. It showed excellent repeatability, with 0.05–0.90% intra-assay variation and 0.02–0.94% inter-assay variation. The 1028 clinical samples were tested using the quadruplex RT-qPCR and a reported reference RT-qPCR. The positivity rates of CCoV, CRV, CPV, and CDV were 9.53%, 0.97%, 25.68%, and 5.06% using the developed assay, and 9.05%, 0.88%, 25.68%, and 4.86% using the reference assay, with agreements higher than 99.32%. Conclusion: The results indicated that a rapid and accurate quadruplex RT-qPCR was developed for the detection and differentiation of CCoV, CRV, CPV, and CDV.
Full article
(This article belongs to the Special Issue Veterinary Microbiology and Diagnostics)
►▼
Show Figures

Figure 1
Open AccessArticle
In Saccharomyces cerevisiae ρ0 Cells, UME6 Contributes to the Activation of ABC Transporter Genes and Pleiotropic Drug Resistance via RPD3 and PDR3
by
Mai Funasaka, Mahiro Ota and Yoichi Yamada
Microbiol. Res. 2024, 15(2), 734-745; https://doi.org/10.3390/microbiolres15020048 - 9 May 2024
Abstract
►▼
Show Figures
In Saccharomyces cerevisiae, the Rpd3L complex includes the histone deacetylase Rpd3 and the DNA binding proteins Ume6 and Ash1 and serves as a transcriptional silencer or enhancer. In S. cerevisiae, the transcription of PDR5, which encodes a major drug efflux
[...] Read more.
In Saccharomyces cerevisiae, the Rpd3L complex includes the histone deacetylase Rpd3 and the DNA binding proteins Ume6 and Ash1 and serves as a transcriptional silencer or enhancer. In S. cerevisiae, the transcription of PDR5, which encodes a major drug efflux pump, and pleiotropic drug resistance (PDR) are hyperactivated by the transcription factor Pdr3 in ρ0/− cells, which lack mitochondrial DNA. We previously showed that RPD3 and UME6 are required for the activation of PDR5 transcription and PDR in S. cerevisiae ρ0 cells. Here, using real-time PCR analysis, we revealed that RPD3 and UME6 are responsible for the activated basal expression of the ABC transporter-encoding genes SNQ2, PDR15, and PDR5 in S. cerevisiae ρ0 cells. Furthermore, using real-time PCR analysis and a spot dilution assay, we found that Ume6 increases the basal expression of PDR5 and PDR15 and induces PDR in a manner dependent on RPD3 and PDR3 in ρ0 cells. This finding may contribute to the elucidation of the relationships between the molecules required for the activation of ABC transporter genes in S. cerevisiae ρ0/− cells and in pathogenic Candida species.
Full article
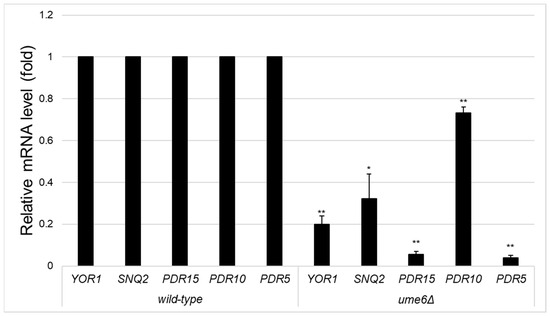
Figure 1
Open AccessArticle
Weissella koreensis KJ, Which Increases Gut Tight Junction Protein Expression, Alleviates TNBS-Induced Colitis by Suppressing Inflammatory Cytokines
by
Kyung-Joo Kim, Hyoleem Lee, Yoon Sin Oh and Se-Eun Jang
Microbiol. Res. 2024, 15(2), 721-733; https://doi.org/10.3390/microbiolres15020047 - 8 May 2024
Abstract
►▼
Show Figures
Inflammatory bowel disease (IBD), a chronic inflammatory disease, results from dysregulation of the immune responses. The IBD prevalence rate was 321.2 per 100,000 people in 2021 and, compared with that in 2006 (200 per 100,000 people), had increased at a rate of +46%.
[...] Read more.
Inflammatory bowel disease (IBD), a chronic inflammatory disease, results from dysregulation of the immune responses. The IBD prevalence rate was 321.2 per 100,000 people in 2021 and, compared with that in 2006 (200 per 100,000 people), had increased at a rate of +46%. Therefore, the development of a safe and new treatment for IBD is urgently needed. Weissella koreensis, a strain of lactic acid bacteria (LABs), was isolated from kimchi and shown to inhibit a pro-inflammatory cytokine, tumor necrosis factor-alpha (TNF-α). Its anti-inflammatory effect was further assessed using a mouse model of colitis induced by 2,4,6-trinitrobenzenesulfonic acid (TNBS). The administration of TNBS significantly increased myeloperoxidase (MPO) expression, macroscopic score, and colonic shortening. Oral administration of W. koreensis KJ suppressed the TNBS-induced response and significantly inhibited the expression of the pro-inflammatory cytokines TNF-α, interleukin (IL)-1β, and IL-6 in the intestinal tissues. In particular, W. koreensis KJ reversed the TNBS-induced decrease in the expression of these tight junction proteins. Therefore, since W. koreensis KJ isolated from kimchi, which increases gut tight junction proteins, attenuating colitis by suppressing inflammatory cytokines, it can be used as a therapeutic candidate for treating colitis such as IBD.
Full article

Figure 1
Open AccessReview
Ice-Nucleating Gut Microbes in Insects: A Scoping Review
by
Matan Shelomi
Microbiol. Res. 2024, 15(2), 708-720; https://doi.org/10.3390/microbiolres15020046 - 30 Apr 2024
Abstract
►▼
Show Figures
(1) Background. At subzero temperatures, water crystallizes on ice nucleation agents. Researchers have identified ice-nucleating microbes (INMs) in insect digestive tracts that can raise the insect’s supercooling point, causing freezing at higher temperatures but slower rates. For freeze-tolerant insects, such gut microbes should
[...] Read more.
(1) Background. At subzero temperatures, water crystallizes on ice nucleation agents. Researchers have identified ice-nucleating microbes (INMs) in insect digestive tracts that can raise the insect’s supercooling point, causing freezing at higher temperatures but slower rates. For freeze-tolerant insects, such gut microbes should allow for slower freezing away from tissues and higher survival rates. For freeze-susceptible insects, however, such microbes could cause a fatal freeze at higher temperatures, and could possibly be used as biocontrol. (2) Methods. A first-ever scoping review was carried out of research on insect-associated INMs, from observational studies attempting to isolate these microbes, to experimental studies applying them and checking for increased mortality. (3) Results. Relatively few research groups have studied insect-associated INMs in any capacity. (4) Conclusions. Several authors hypothesized that such microbes are probably abundant, and their contribution to ice nucleation activity in insects is under-reported. Biocontrol assays using ice-nucleating microbes showed promise, but a risk to non-target organisms has been experimentally confirmed. Future surveys of insect–microbe interactions using molecular tools are likely to reveal new examples, if not new microbe species capable of ice nucleation.
Full article
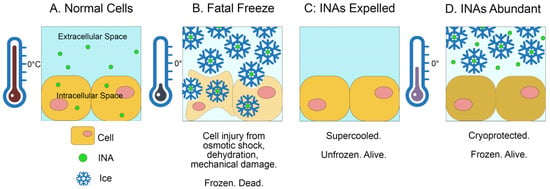
Figure 1
Open AccessArticle
Genetic Diversity of Plasmodium vivax Surface Ookinete Protein Pvs25 and Host Genes in Individuals Living along the Thai–Myanmar Border and Their Relationships with Parasite Density
by
Abdifatah Abdullahi Jalei, Wanna Chaijaroenkul and Kesara Na-Bangchang
Microbiol. Res. 2024, 15(2), 693-707; https://doi.org/10.3390/microbiolres15020045 - 30 Apr 2024
Abstract
►▼
Show Figures
Plasmodium vivax (Pv) accounts for over 50% of malaria cases in Latin America and Asia. Despite a significant reduction in Pv transmission in Thailand, the parasite remains endemic to the border areas. This study aimed to investigate the genetic diversity of
[...] Read more.
Plasmodium vivax (Pv) accounts for over 50% of malaria cases in Latin America and Asia. Despite a significant reduction in Pv transmission in Thailand, the parasite remains endemic to the border areas. This study aimed to investigate the genetic diversity of the parasites and the host factors, as well as their relation to parasite density in Pvisolates, along the Thai–Myanmar border. Genetic variations in Pv markers, specifically the ookinete surface protein Pvs25, and host genes, including Toll-like receptor 6 (TLR6), TLR9, TIR Domain-containing adaptor protein (TIRAP), Toll-interacting protein (TOLLIP), Duffy antigen receptor for chemokines (DARC), and intercellular adhesion molecule 1 (ICAM-1), were investigated using polymerase chain reaction (PCR) with restriction fragment length polymorphism (RFLP). A total of 548 PCR-positive Pv samples collected from Tak and Kanchanaburi provinces during two periods (2006–2007 and 2014–2016) were included in the study. Pvs25 exhibited four haplotypes, with H1 (EGTKV) being the most prevalent in both provinces. Kanchanaburi isolates exhibited greater genetic diversity than Tak isolates. No significant deviations from neutrality were observed for Pvs25 in either area. ICAM-1 and TOLLIP s3750920 heterozygous carriers had greater median parasite densities than homozygous mutants. The TLR9 rs187084 T genotype had a significantly higher parasite density than the non-T genotype. The findings underscore the significant association between the rs3750920 C/T, rs5498 A/G, and rs187084 T genotypes and high parasite density in patients infected with Pv, highlighting their potentially critical role in malaria susceptibility.
Full article
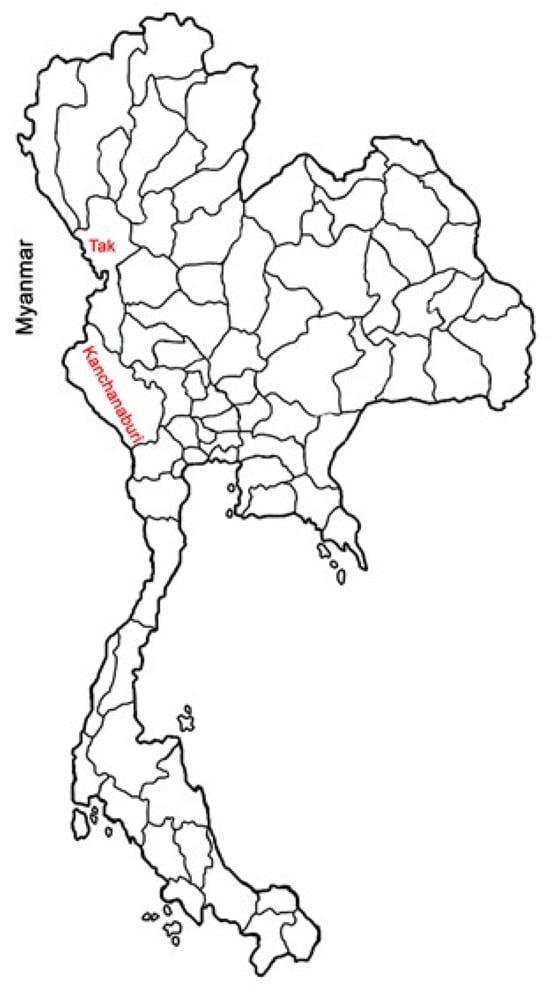
Figure 1
Open AccessReview
An Update on Zika Virus Vaccine Development and New Research Approaches
by
Angie Lizeth Buitrago-Pabón, Salvador Ruiz-Sáenz, Alicia Jiménez-Alberto, Gerardo Aparicio-Ozores, Juan Arturo Castelán-Vega and Rosa María Ribas-Aparicio
Microbiol. Res. 2024, 15(2), 667-692; https://doi.org/10.3390/microbiolres15020044 - 29 Apr 2024
Abstract
►▼
Show Figures
Zika virus (ZIKV) is an emerging flavivirus that represents significant public health challenges, particularly in the Americas, and is a substantial risk to other parts of the world due to its rapid expansion and its established association with neurological disorders, including Guillain–Barré syndrome
[...] Read more.
Zika virus (ZIKV) is an emerging flavivirus that represents significant public health challenges, particularly in the Americas, and is a substantial risk to other parts of the world due to its rapid expansion and its established association with neurological disorders, including Guillain–Barré syndrome and an intrauterine fetal infection that can cause microcephaly, blindness, and other congenital neurological complications. To date, no vaccine to prevent ZIKV infections has been approved. Therefore, developing a safe and effective vaccine against this virus is a global health priority. This review analyzes the ZIKV outbreaks, as well as associated neurological complications, its genome, and immunological responses. The current vaccines in development have reported results from preclinical and clinical trials about novel approaches to obtain safer and more effective vaccines and the challenges faced by ZIKV vaccine development.
Full article
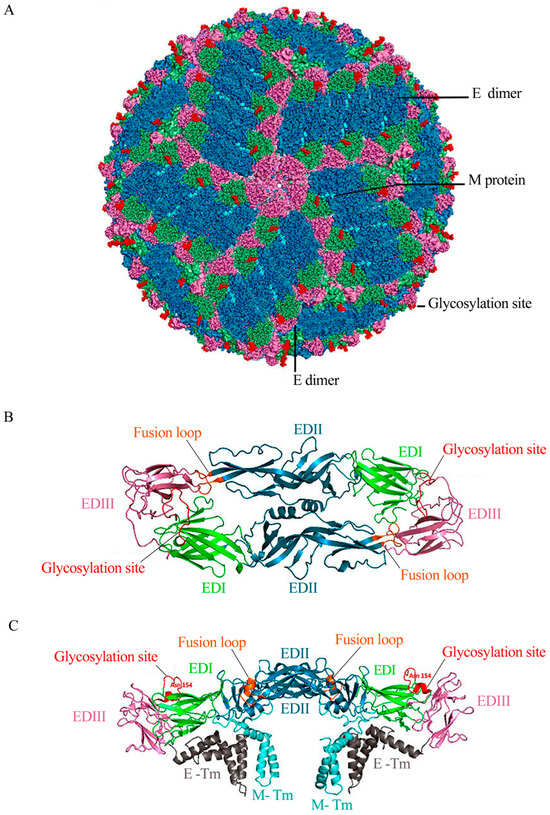
Figure 1
Open AccessArticle
Exposure of Dairy Cows to Coxiella burnetii in Greece: Surveillance Results and Association of Bacterial Presence with Environmental Variables
by
George Valiakos, Ioannis Gouvias, Marios Lysitsas, Ilias Bouzalas, Stefania Tampach, Eleni Malissiova, Alexis Giannakopoulos, Constantina N. Tsokana, Dimitrios Vourvidis, Anna Kyrma and Charalambos Billinis
Microbiol. Res. 2024, 15(2), 655-666; https://doi.org/10.3390/microbiolres15020043 - 25 Apr 2024
Abstract
►▼
Show Figures
The exposure of dairy cows to Coxiella burnetii using molecular and serological techniques was investigated in this study. Bulk tank milk and serum samples were collected from various farms in Greece (mainly northern Greece). DNA extraction was performed on milk samples, and qPCR
[...] Read more.
The exposure of dairy cows to Coxiella burnetii using molecular and serological techniques was investigated in this study. Bulk tank milk and serum samples were collected from various farms in Greece (mainly northern Greece). DNA extraction was performed on milk samples, and qPCR targeting the IS1111 insertion sequence was performed to detect bacterial pathogens. An ELISA was used to detect specific antibodies in bulk milk and individual serum samples. Data on farms were collected in the field using handheld Global Positioning System Garmin units. The collected data were analyzed using an Ecological Niche Model within the framework of a geographic information system. The results indicated that in more than half of the dairy farms (35/60, 58.3%), C. burnetii is present in milk. Specific antibodies were also detected in almost all milk samples (57/60, 95.0%). At least one seropositive animal was identified using ELISA in the majority of the examined farms (25/28, 89.3%). C. burnetii PCR-positive farms were located in the low-altitude zone with a mean value of 97 m above sea level (range: 2–681). The environmental variable with the highest gain when used in isolation is precipitation in the wettest quarter (28.3% contribution), which therefore appears to have the most useful information by itself. The environmental variable that decreases the gain the most when omitted is the minimal temperature of the coldest month (6.9% contribution). The analysis demonstrated that a mild climate with low precipitation favors a positive status. The exposure of dairy cattle farms to C. burnetii is massive, raising significant concerns regarding livestock production and public health implications.
Full article
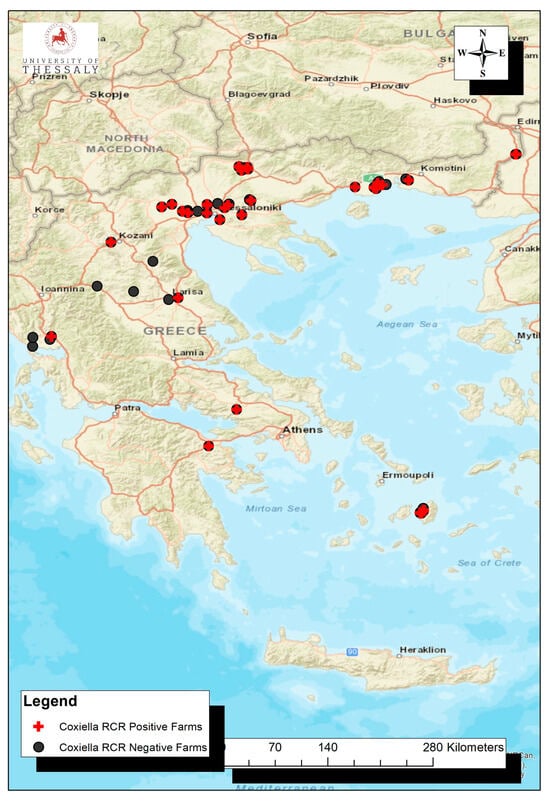
Figure 1
Open AccessReview
Methanotrophy: A Biological Method to Mitigate Global Methane Emission
by
Anju Rani, Aarushi Pundir, Medhashree Verma, Samiksha Joshi, Geeta Verma, Snežana Andjelković, Snežana Babić, Jasmina Milenković and Debasis Mitra
Microbiol. Res. 2024, 15(2), 634-654; https://doi.org/10.3390/microbiolres15020042 - 25 Apr 2024
Abstract
►▼
Show Figures
Methanotrophy is a biological process that effectively reduces global methane emissions by utilizing microorganisms that can utilize methane as a source of energy under both oxic and anoxic conditions, using a variety of different electron acceptors. Methanotrophic microbes, which utilize methane as their
[...] Read more.
Methanotrophy is a biological process that effectively reduces global methane emissions by utilizing microorganisms that can utilize methane as a source of energy under both oxic and anoxic conditions, using a variety of different electron acceptors. Methanotrophic microbes, which utilize methane as their primary source of carbon and energy, are microorganisms found in various environments, such as soil, sediments, freshwater, and marine ecosystems. These microbes play a significant role in the global carbon cycle by consuming methane, a potent greenhouse gas, and converting it into carbon dioxide, which is less harmful. However, methane is known to be the primary contributor to ozone formation and is considered a major greenhouse gas. Methane alone contributes to 30% of global warming; its emissions increased by over 32% over the last three decades and thus affect humans, animals, and vegetation adversely. There are different sources of methane emissions, like agricultural activities, wastewater management, landfills, coal mining, wetlands, and certain industrial processes. In view of the adverse effects of methane, urgent measures are required to reduce emissions. Methanotrophs have attracted attention as multifunctional bacteria with potential applications in biological methane mitigation and environmental bioremediation. Methanotrophs utilize methane as a carbon and energy source and play significant roles in biogeochemical cycles by oxidizing methane, which is coupled to the reduction of various electron acceptors. Methanotrophy, a natural process that converts methane into carbon dioxide, presents a promising solution to mitigate global methane emissions and reduce their impact on climate change. Nonetheless, additional research is necessary to enhance and expand these approaches for extensive use. In this review, we summarize the key sources of methane, mitigation strategies, microbial aspects, and the application of methanotrophs in global methane sinks with increasing anthropogenic methane emissions.
Full article
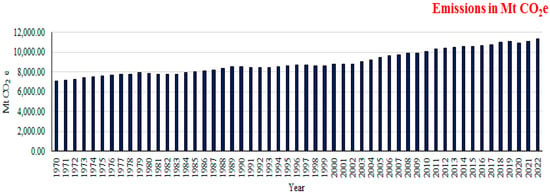
Figure 1
Open AccessCase Report
First Molecular Detection and Characterization of Fowl Aviadenovirus Serotype 11 from Broiler Chickens in Chile
by
Leandro Cádiz, Miguel Guzmán, Fernando Navarrete, Paulina Torres and Hector Hidalgo
Microbiol. Res. 2024, 15(2), 626-633; https://doi.org/10.3390/microbiolres15020041 - 23 Apr 2024
Abstract
Fowl aviadenovirus (FAdV) is a member of the Aviadenovirus genus within the Adenoviridae family. FAdVs are divided into five species based on genomic differences: Fowl aviadenovirus A to Fowl aviadenovirus E (FAdV-A to FAdV-E). They are classified into twelve serotypes (FAdV-1 to FAdV-8a
[...] Read more.
Fowl aviadenovirus (FAdV) is a member of the Aviadenovirus genus within the Adenoviridae family. FAdVs are divided into five species based on genomic differences: Fowl aviadenovirus A to Fowl aviadenovirus E (FAdV-A to FAdV-E). They are classified into twelve serotypes (FAdV-1 to FAdV-8a and FAdV-8b to FAdV-11) through cross-neutralization tests. FAdVs are mainly associated with hepatitis hydropericardium syndrome (HHS), adenoviral gizzard erosion (AGE), and inclusion body hepatitis (IBH). The serotypes commonly involved in IBH are FAdV-2, FAdV-11, FAdV-8a, and FAdV-8b. IBH causes significant economic losses in the poultry industry, mainly due to high mortality, reduced productivity, and immunosuppression. This is the first case report on IBH in Chile caused—according to post-mortem findings, molecular analysis, sequencing, and phylogenetic analysis—by FAdV-11. Since the serotype had not previously been reported in Chile, continued monitoring of IBH cases is required to determine the serotype of the circulating FAdVs and adapt preventative vaccination programs.
Full article
(This article belongs to the Special Issue Evolution of Viral Virulence)
►▼
Show Figures
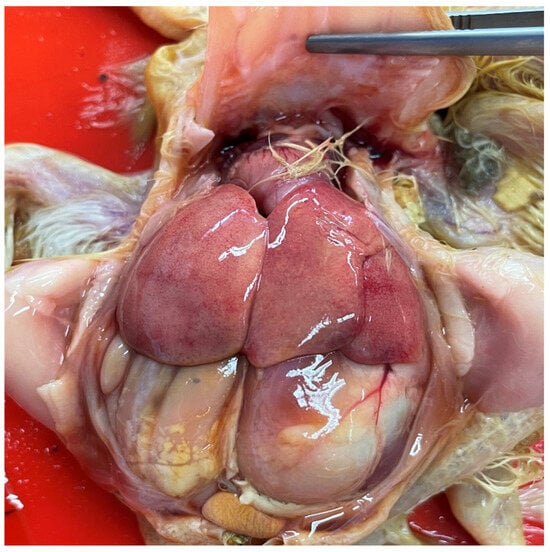
Figure 1
Open AccessArticle
Ingested Microplastics Can Act as Microbial Vectors of Ichthyofauna
by
Abdulhusein Jawdhari, György Deák, Dan Florin Mihăilescu, Nicolai Crăciun, Andrea Cristina Staicu, Ioana Stanca, Derniza Cozorici, Sergiu Fendrihan, Cristian-Emilian Pop and Maria Mernea
Microbiol. Res. 2024, 15(2), 614-625; https://doi.org/10.3390/microbiolres15020040 - 23 Apr 2024
Abstract
Microplastics (plastic particles < 5 mm) are ubiquitous pollutants that have the ability to carry microbiota, including pathogens. Microbial adhesion is usually a sign of pathogenicity; thus, we investigated the adherent microbiota found on 4 mm nylon strips, which were ingested and excreted
[...] Read more.
Microplastics (plastic particles < 5 mm) are ubiquitous pollutants that have the ability to carry microbiota, including pathogens. Microbial adhesion is usually a sign of pathogenicity; thus, we investigated the adherent microbiota found on 4 mm nylon strips, which were ingested and excreted by wild fish specimens. Retention times were recorded and the polymer analysis of the excreted samples was performed, which showed no signs of degradation, nor did their controls, represented by the nylon strips submerged in the same water tanks. Both the ingested samples and controls presented pathogens in large quantities. Following Matrix-Assisted Laser Desorption/Ionization Time-Of-Flight identification, the dominant genus was represented by Aeromonas, revealing the fact that nylon microplastics can serve as undegradable physical carriers for this pathogen, among others, in the aquatic environment.
Full article
(This article belongs to the Special Issue Zoonotic Bacteria: Infection, Pathogenesis and Drugs)
►▼
Show Figures
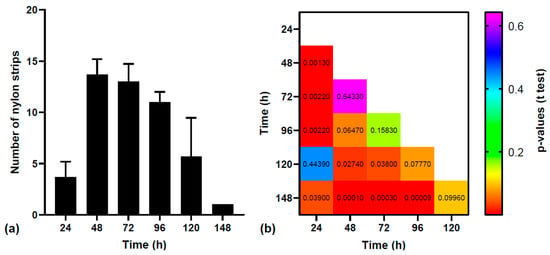
Figure 1
Open AccessArticle
Correlation between Aerosol Particulates, Carcass Dirtiness, and Hygiene Indicators of Bovine Carcasses in the Abattoir Environment: Results of a Study in Italy
by
Beniamino T. Cenci-Goga, Emma Tedeschini, Egidia Costanzi, Margherita Maranesi, Musafiri Karama, Saeed El-Ashram, Cristina Saraiva, Juan García-Díez, Massimo Zerani, Ebtesam M. Al-Olayan and Luca Grispoldi
Microbiol. Res. 2024, 15(2), 598-613; https://doi.org/10.3390/microbiolres15020039 - 22 Apr 2024
Abstract
The objective of this study was to demonstrate the possible correlation of visible carcass contamination and abattoir aerosol quality with microbial hygiene criteria. A total of 279 bovine carcasses were analyzed on 23 different working days. The aerobic colony count and total coliforms
[...] Read more.
The objective of this study was to demonstrate the possible correlation of visible carcass contamination and abattoir aerosol quality with microbial hygiene criteria. A total of 279 bovine carcasses were analyzed on 23 different working days. The aerobic colony count and total coliforms on the carcasses were calculated together with the presence of Escherichia coli. To determine the visible contamination of carcasses, we used a 100 cm2 sheet of transparent, adhesive plastic material, applied to the side of the carcass, to collect all the particles, which were then counted against both black and white backgrounds. The daily particulate index in the abattoir aerosol was determined using an air sampler device. The results showed that aerobic colony counts, which ranged from 1.41 to 2.40 log cfu cm−2, total coliforms (from 0.00 to 0.73 log cfu cm−2), and E. coli presence (from 0.00% to 60% of the sampled carcasses per day) are not correlated with the carcasses’ visual dirtiness or the aerosol quality. The factor analysis showed a correlation between the three groups of variables investigated: group 1, representing “aerosol quality”, group 2, representing the “microbiology of the carcass”, and group 3, the “visual dirtiness of the carcass”. Thus, even though microbiology analysis is useful in diagnosing the microorganisms which the official veterinarian is unable to detect during the post-mortem inspection, it is ineffective in evaluating slaughtering procedures. Aerosol monitoring and the visual classification of carcass dirtiness, instead, could provide good indications of the slaughtering process and the quality of the abattoir environment, and guarantee control of manufacturing practices, protecting both animals’ and operators’ health.
Full article
(This article belongs to the Special Issue Molecular Microbiology Underlying Foodborne Viruses)
►▼
Show Figures
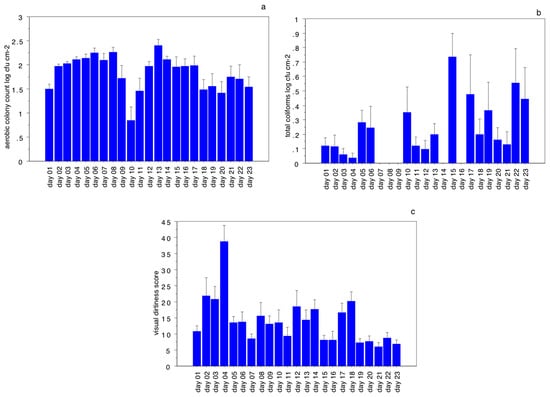
Figure 1
Open AccessArticle
Antiviral Activity of Flavonoids from Bauhinia holophylla Leaves against Zika virus
by
Rodrigo Michelini de Oliveira Thomasi, Thaiz Rodrigues Teixeira, Gabriela Francine Martins Lopes, Simony Carvalho Mendonça, Brendo Araujo Gomes, Suzana Guimarães Leitão, Tiago Alves de Oliveira, Sara Thamires Dias da Fonseca, Alex Gutterres Taranto, Jaqueline Maria Siqueira Ferreira, Luciana Alves Rodrigues dos Santos Lima and Ana Hortência Fonsêca Castro
Microbiol. Res. 2024, 15(2), 582-597; https://doi.org/10.3390/microbiolres15020038 - 21 Apr 2024
Abstract
►▼
Show Figures
Zika virus (ZIKV) is involved in the etiology of serious nervous system pathologies. Currently, there are no specific and effective vaccines or antiviral drugs to prevent the diseases caused by ZIKV. This study aimed to assess the activity of flavonoids present in crude
[...] Read more.
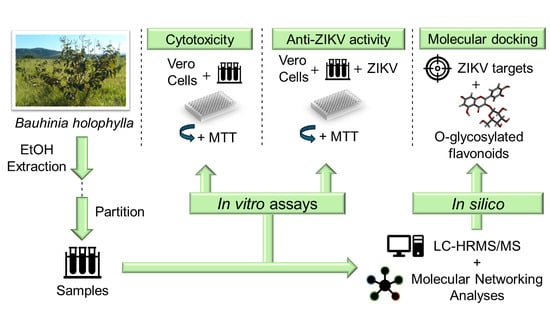
Zika virus (ZIKV) is involved in the etiology of serious nervous system pathologies. Currently, there are no specific and effective vaccines or antiviral drugs to prevent the diseases caused by ZIKV. This study aimed to assess the activity of flavonoids present in crude hydroethanolic extract (CHE) and fractions obtained from B. holophylla leaves against ZIKV. O-glycosylated flavonoids were characterized by high-performance liquid chromatography coupled with high-resolution mass spectrometry (LC-HRMS/MS). The cytotoxic concentration and the effective concentration for 50% of the cells (CC50 and EC50, respectively) were determined, and the selectivity index (SI) was calculated. Molecular networks were constructed based on the chemical composition of the samples and global antiviral activity data using the Global Natural Products Social Molecular Networking (GNPS) platform. Protein–ligand docking was performed in the NS2B-NS3 protease, NS3 helicase, and NS5 methyltransferase of the ZIKV. CHE showed greater antiviral activity at a multiplicity of infection (MOI) of 1.0, with an EC50 of 11.93 µg/mL, SI = 13.38, and reduced cytopathic effects. Molecular networks indicated that O-glycosylated flavonoids are responsible for the activity against ZIKV, being quercetin-O-deoxyhexoside more selective and effective. Molecular docking confirmed the inhibitory activity of quercetin-O-deoxyhexoside, which showed an affinity for the tested targets, especially for NS2B-NS3 protease. The results showed that B. holophylla has flavonoids with potential for future therapeutic applications against ZIKV.
Full article
Graphical abstract
Open AccessArticle
Molecular Biological Methods to Assess Different Botrytis cinerea Strains on Grapes
by
Louis Backmann, Katharina Schmidtmann, Pascal Wegmann-Herr, Andreas Jürgens and Maren Scharfenberger-Schmeer
Microbiol. Res. 2024, 15(2), 567-581; https://doi.org/10.3390/microbiolres15020037 - 19 Apr 2024
Abstract
►▼
Show Figures
Botrytis cinerea is a well-known pathogen that can be challenging to control in crops, such as wine grapes. To adapt to the increasing problems of climate change and strain resistance, it is important to find new methods to detect Botrytis cinerea and differentiate
[...] Read more.
Botrytis cinerea is a well-known pathogen that can be challenging to control in crops, such as wine grapes. To adapt to the increasing problems of climate change and strain resistance, it is important to find new methods to detect Botrytis cinerea and differentiate strains. These methods include strain differentiation and classification by simple sequence repeats (SSRs) and early detection of the fungus by qPCR. Various strains were analysed using SSR markers and either agarose gel electrophoresis or capillary sequencing via PCR. A sensitive qPCR method was refined to achieve an early detection method for the pathogen. The results demonstrate promising ways to distinguish between strains using both agarose gel electrophoresis and capillary sequencing as well as to detect infection before it becomes visible on grapes. This can be used to further understand and analyse different Botrytis cinerea strain characteristics such as laccase activity, regional or annual effects. The early detection method can be used to better prepare growers for an impending infection so that targeted efforts can be made.
Full article
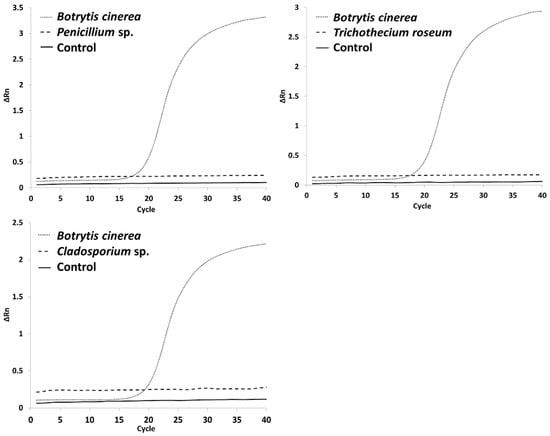
Figure 1
Open AccessArticle
Altered Cytostructure and Lignolytic Enzymes of Ganoderma boninense in Response to Phenolic Compounds
by
Yasmeen Siddiqui and Daarshini Ganapathy
Microbiol. Res. 2024, 15(2), 550-566; https://doi.org/10.3390/microbiolres15020036 - 16 Apr 2024
Abstract
►▼
Show Figures
Ganoderma boninense is a white-rot fungus that causes basal stem rot (BSR) disease in the oil palm. Potential natural inhibitors, such as gallic acid, thymol, propolis, and carvacrol, were assessed for their antagonistic effects against G. boninense. These naturally occurring phenolic compounds
[...] Read more.
Ganoderma boninense is a white-rot fungus that causes basal stem rot (BSR) disease in the oil palm. Potential natural inhibitors, such as gallic acid, thymol, propolis, and carvacrol, were assessed for their antagonistic effects against G. boninense. These naturally occurring phenolic compounds have also been utilised to inhibit hydrolytic and ligninolytic enzymes produced by the pathogen. Mycelial inhibition was dose-dependent in the presence of different concentrations of phenolic compounds, including, for example, in cellulase enzyme inhibition (GA mg/mL = 94%, THY 0.25 mg/mL = 90%, PRO 3.5 mg/mL = 92.5%, and CARV 0.15 mg/mL = 90.3%). A significant difference was observed revealing that gallic acid had the greatest inhibitory effect on the secretion of hydrolytic and ligninolytic enzymes, especially at 40 mM GA (cellulase = 0.337 U/mL, amylase = 0.3314 U/mL, xylanase = 0.211 U/mL, laccase = 0.4885 U/mL, lignin peroxidase = 0.218 U/mL, and manganese peroxidase = 0.386 U/mL). The growth and secretion of enzymes (inhibitory action) are inversely proportional to the concentration of phenolic compounds. Phenolic compounds have a greater potential as inhibitory agents and suppress the production of hydrolytic and ligninolytic enzymes. The selected phenolic compounds were evaluated for their ability to alter the morphology and integrity of G. boninense mycelia. The reduction in cell viability of G. boninense has been explained by research on morphological disruption, such as branching patterns, hyphal length, and rigidity of fungal cells, which eventually interrupt the secretion of enzymes. These studies highlight the efficacy of phenolic compounds in treating Ganoderma. In addition, these findings proved that naturally occurring phenolic compounds could be a substitute for chemical controls and other synthetic fungicides to eradicate the occurrence of BSR in oil palms, thus avoiding a situation that is difficult to overcome.
Full article

Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Diseases, IJMS, Microbiology Research, Pathogens, Vaccines
Advances in Human Pathogen Control—a 21st Century Challenge 2.0
Topic Editors: Jorge H. Leitão, Nitin Amdare, Joana R FelicianoDeadline: 30 June 2024
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024
Topic in
Gastroenterology Insights, Infectious Disease Reports, Life, Microbiology Research, Microorganisms
High-Throughput Analyses as a Multi-Faceted Approach for Characterizing the Human Microbiota
Topic Editors: Simone Filardo, Rosa Sessa, Andrea Carolina EntrocassiDeadline: 30 October 2024

Conferences
Special Issues
Special Issue in
Microbiology Research
Zoonotic Bacteria: Infection, Pathogenesis and Drugs
Guest Editors: Yang Wang, Jiazhang QiuDeadline: 31 May 2024
Special Issue in
Microbiology Research
Benefits and Applications of Probiotics in Aquaculture
Guest Editors: Hany M.R. Abdel-Latif, Sevdan YilmazDeadline: 30 June 2024
Special Issue in
Microbiology Research
Antifungal Activities of Plant Extracts
Guest Editor: Julliana Ribeiro Alves SantosDeadline: 25 July 2024
Special Issue in
Microbiology Research
Veterinary Microbiology and Diagnostics
Guest Editor: Seyed Ali GhorashiDeadline: 31 August 2024
Topical Collections
Topical Collection in
Microbiology Research
Microbiology and Technology of Fermented Foods
Collection Editor: Salam A. Ibrahim
Topical Collection in
Microbiology Research
Public Health and Quality Aspects Related to Animal Productions
Collection Editors: Beniamino T. Cenci-Goga, Massimo Zerani, Luca Grispoldi
Topical Collection in
Microbiology Research
Microorganisms and Their Incredible Potential to Face Societal Challenges
Collection Editor: Mireille Fouillaud








